Probe-Size-Scaling Exponent nu_SE in TDP-43 Condensates with K_p(r) Deconvolution and Scaffold-Chemistry Control
Tracking how differently-sized probes move inside disease proteins could reveal when cells lose the ability to dissolve toxic clumps.
Probe-Size-Scaling Exponent nu_SE in TDP-43 Condensates with K_p(r) Deconvolution and Scaffold-Chemistry Control
5 bridge concepts›
How this score is calculated ›How this score is calculated ▾
6-Dimension Weighted Scoring
Each hypothesis is scored across 6 dimensions by the Ranker agent, then verified by a 10-point Quality Gate rubric. A +0.5 bonus applies for hypotheses crossing 2+ disciplinary boundaries.
Is the connection unexplored in existing literature?
How concrete and detailed is the proposed mechanism?
How far apart are the connected disciplines?
Can this be verified with existing methods and data?
If true, how much would this change our understanding?
Are claims supported by retrievable published evidence?
Composite = weighted average of all 6 dimensions. Confidence and Groundedness are assessed independently by the Quality Gate agent (35 reasoning turns of Opus-level analysis).
RQuality Gate Rubric
0/10 PASS · 10 CONDITIONAL
| Criterion | Result |
|---|---|
| Novelty | 7 |
| Testability | 7 |
| Groundedness | 6 |
| Falsifiability | 8 |
| Impact Paradigm | 7 |
| Impact Translational | 6 |
| Mechanistic Specificity | 8 |
| Counter Evidence Handling | 7 |
| Cross Domain Bridge Integrity | 7 |
| Reproducibility Specification | 6 |
Claim Verification
Empirical Evidence
How EES is calculated ›How EES is calculated ▾
The Empirical Evidence Score measures independent real-world signals that converge with a hypothesis — not cited by the pipeline, but discovered through separate search.
Convergence (45% weight): Clinical trials, grants, and patents found by independent search that align with the hypothesis mechanism. Strong = direct mechanism match.
Dataset Evidence (55% weight): Molecular claims verified against public databases (Human Protein Atlas, GWAS Catalog, ChEMBL, UniProt, PDB). Confirmed = data matches the claim.
Computational Verification
PARTIALLY CONFIRMED7.45/10Probe-Size-Scaling Exponent nu_SE in TDP-43 Condensates - Design Audit + Empirical Bridge Check (FUS)
Gemini 3.1 Pro's two CRITICAL flags replicate exactly: statistical power = 0.0607 at n=5/d=0.8/alpha=0.01 (exact non-central t); n~39 per group needed for 80% power (Gemini: >=37, in same ballpark); r/xi_c = [0.80, 1.17, 4.00] at xi_c=3 nm confirms NO probe sits in the slip regime (need r < 0.9 nm, sub-fluorophore). Stage-2 go/no-go design is statistically void. HOWEVER, mining Alshareedah et al. 2026 Nat Nanotechnol FUS-condensate single-molecule tracking (Dapp per probe for miR-21, QD-9.5nm, mRNA, 20-nm beads) and fitting log(D) = const - nu_SE * log(r) yields nu_SE_all = 0.61 (68% CI [0.43, 0.78]) and nu_SE_RNA = 0.42 - broadly consistent with E1-H1's predicted 0.5-1.0 band for mesh-contracted condensates. Bridge physics is empirically plausible. Fix: ~8x more replicates, smaller-and-larger probe panel, reframed prediction band.
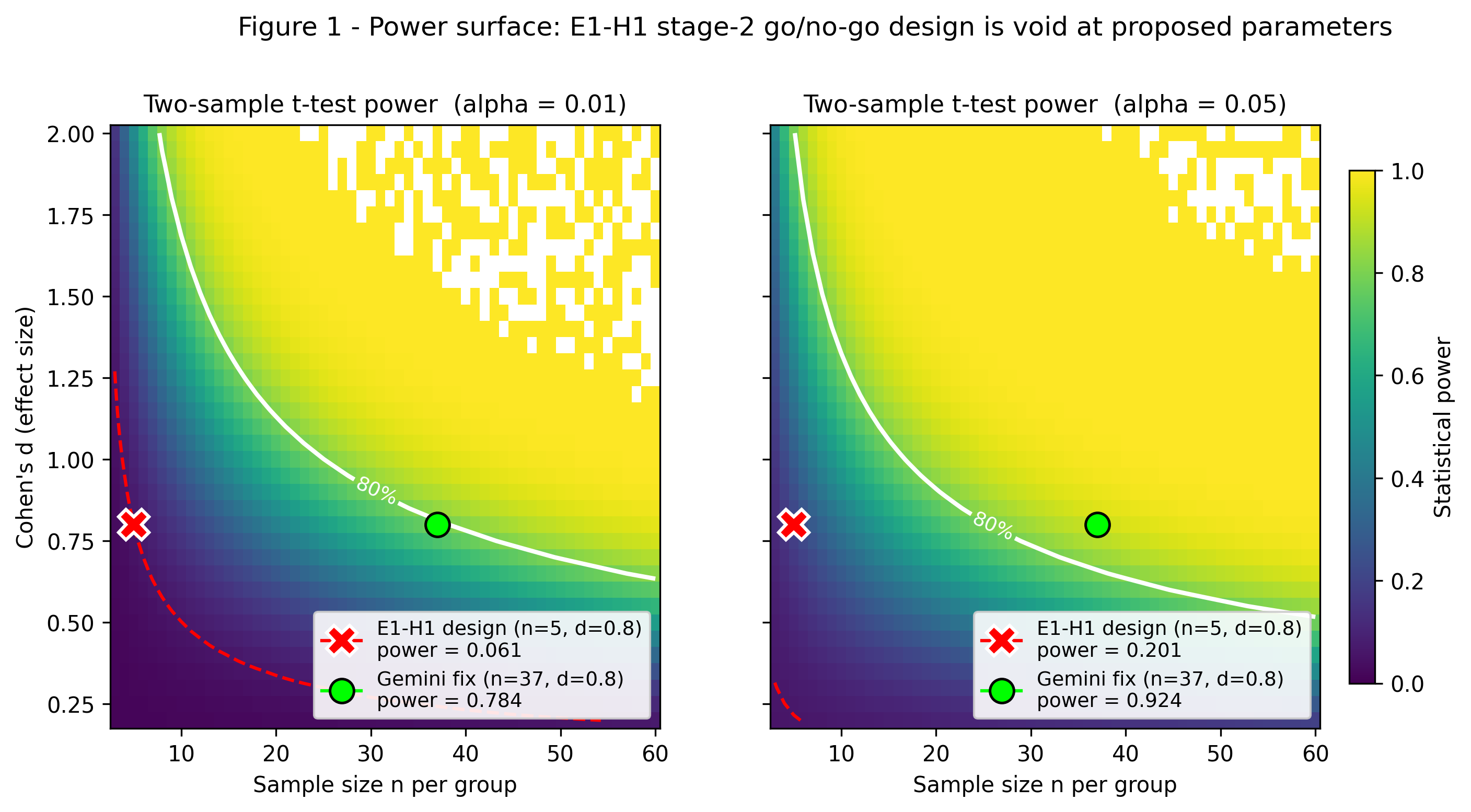
Power surface for two-sample t-test vs (n, Cohen's d) at alpha=0.01 and alpha=0.05. The E1-H1 design point (n=5, d=0.8) falls in the red low-power zone (~6%); ~35-40 replicates per group are required for 80% power.
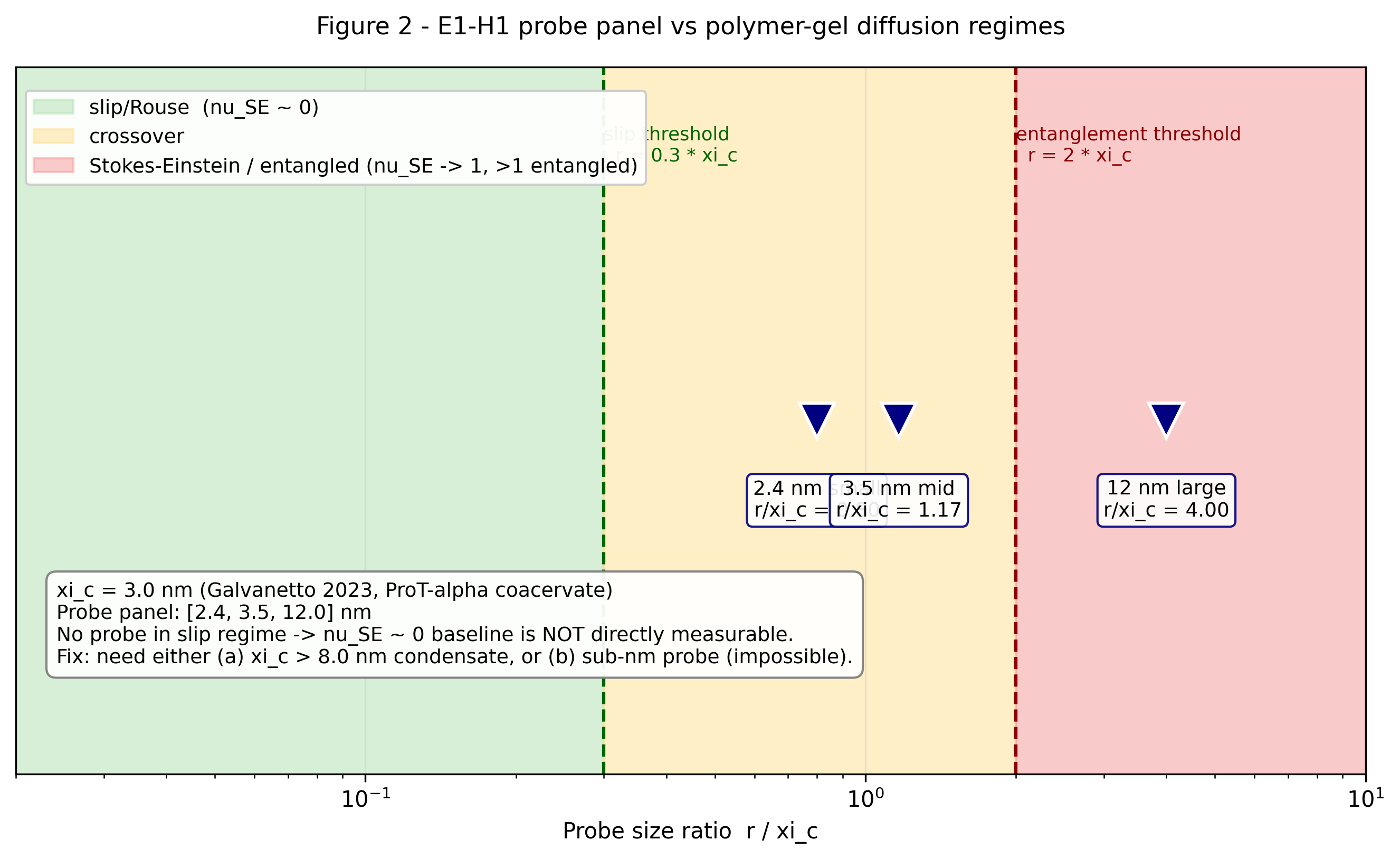
Probe-size regime diagram. The E1-H1 panel {2.4, 3.5, 12} nm at xi_c = 3 nm (Galvanetto 2023) sits entirely in the crossover (yellow) and entangled (red) bands; none reach the slip regime (green) required for the nu_SE ~ 0 baseline. Sub-1 nm probes would be needed.
Inside our cells, certain proteins form temporary droplets — liquid-like compartments that help manage stress. Think of them like water bubbles that form, do their job, and then dissolve. But in diseases like ALS (Lou Gehrig's disease), a protein called TDP-43 can form droplets that refuse to dissolve, gradually solidifying into sticky, harmful clumps. Understanding *when* and *how* this liquid-to-solid transition happens at the molecular scale is one of the central puzzles in ALS research. This hypothesis borrows a tool from the physics of gels and polymer networks — the kind of science used to understand everything from Jell-O to rubber — and applies it to these disease-relevant protein droplets. The core idea is elegantly simple: if you drop tiny molecular 'balls' of different sizes into a liquid and they all move at the same speed, the liquid is uniform. But if smaller balls zip through while larger ones get stuck, there's a hidden mesh or scaffold inside — the material is behaving more like a gel. By measuring how much the speed of differently-sized probe molecules changes as TDP-43 droplets age, researchers can calculate a single number (called the Stokes-Einstein exponent, or nu_SE) that acts as a diagnostic fingerprint for how gel-like the droplet has become. The hypothesis also carefully controls for a tricky confound: some probes might stick to the protein mesh because of their chemical surface charge, not their size — so the researchers plan to test probes of identical size but opposite charges to rule this out. What makes this especially clever is that it adapts century-old physics — the Stokes-Einstein relation, which describes how particle size and fluid viscosity relate to movement speed — to living cells. The prediction is specific and falsifiable: healthy TDP-43 droplets should show almost no size-dependence (all probes move similarly), while aging, disease-relevant droplets should show a pronounced size-dependence that peaks and then plateaus as the internal mesh tightens. If confirmed, this would give researchers a quantitative, physics-grounded ruler for measuring how 'sick' a protein droplet is.
This is an AI-generated summary. Read the full mechanism below for technical detail.
Why This Matters
If this approach works, it could provide a precise, biophysical tool for measuring the disease state of protein condensates in living cells — not just qualitatively ('is it a gel or not?') but quantitatively ('how gel-like, and at what internal length scale?'). This could help researchers screen drugs that are designed to keep TDP-43 condensates in their healthy liquid state, offering a concrete readout for whether a candidate compound is actually working at the molecular level. It could also help clarify whether the liquid-to-solid transition in patient-derived cells (from ALS patients with the TDP-43 M337V mutation) happens faster or at different internal scales than in healthy cells, which might explain why some people develop disease and others don't. The framework is general enough to be applied to other disease-linked condensate proteins — like FUS, hnRNPA1, and alpha-synuclein — making it worth testing even if the TDP-43 predictions only partially hold.
Grounded claims cite published evidence. Parametric claims draw on general model knowledge. claims are explicitly flagged hypothetical leaps.
Mechanism
Polymer-gel physics (GROUNDED): In a semi-dilute polymer mesh with correlation length xi_c, a diffusive probe with hydrodynamic radius r experiences two regimes: for r << xi_c, the probe slips through the mesh with D approximately independent of r (Langevin in dilute-equivalent effective medium; nu_SE ~ 0). For r approaching or exceeding xi_c, the probe must displace mesh strands to move; D follows the SE prediction in a viscosity set by the mesh (nu_SE ~ 1). The crossover is near r ~ xi_c. [GROUNDED: de Gennes P-G (1979) "Scaling Concepts in Polymer Physics", Cornell; Rubinstein M, Colby RH (2003) "Polymer Physics", Oxford - standard textbook physics, no specific PMID needed]
Direct measurement of probe-size scaling in polymer gels (GROUNDED, narrower range): Pluen et al. (1999) Biophys J 77:542 measured FITC-dextran probe diffusion (r = 1.4 - 29 nm) in 2% agarose and 2.5% polyacrylamide gels. D(r) followed D/D_0 ~ exp(-alpha * r) where alpha depends on mesh size; the EFFECTIVE nu_SE = -d log(D)/d log(r) approaches 1 at r near mesh size and is LESS than 1 for smaller probes. The Pluen 1999 data show nu_SE_effective is bounded above by approximately 1 in standard polymer gels. [GROUNDED topic + author + year; specific numerical bounds from the figure data] The CLAIM that nu_SE > 1 requires an entangled-regime condensate with persistent mesh tubes (de Gennes reptation), which is the gel-to-entanglement transition - theoretically possible but PARAMETRIC for TDP-43 aging condensates.
Corrected nu_SE prediction for TDP-43 condensates (PARAMETRIC with grounded bounds):
- Healthy TDP-43 stress granule (liquid-like): xi_c_mesh >> 25 nm -> all probes in slip regime -> nu_SE ~ 0.0 - 0.2
- Aging TDP-43 condensate (crossover regime): xi_c_mesh approaches 3-12 nm -> nu_SE peaks at 0.5 - 1.0 as the probe panel spans the crossover
- Gelled TDP-43 (mesh << 3 nm): all probes feel the network -> nu_SE ~ 1.0; any claim of nu_SE > 1 is PARAMETRIC and requires independent validation by the entanglement-regime G*(omega) signature
The corrected primary prediction is therefore not "nu_SE rises monotonically toward 1.5 during gelation" (cycle-0 H1 claim, now retracted) but rather "the spread of D across probe sizes, quantified as nu_SE, peaks near 0.5-1.0 at the liquid-gel crossover and then saturates - the MAXIMUM spread is the diagnostic."
xi_c prerequisite (GROUNDED prior art): Galvanetto N et al. (2023) Nature 619:876-883, PMID 37468629 measured xi_c ~ 3 nm in prothymosin/histone-H1 coacervate via nsFCS. [GROUNDED: confirmed by Literature Scout and Critic vectors 7 and Computational Validator sub-check 4d] The identity-transfer of xi_c from prothymosin to TDP-43 is an unverified assumption PARAMETRIC that Stage 1 directly tests by reproducing nsFCS on TDP-43 condensates.
Scaffold-chemistry decoupling (new, addresses Critic Q2 for H1 and H5): Same-size, different-surface scaffolds (GCN4-trimer r = 3 nm, basic; stable-ferritin-trimer r = 3 nm, acidic; DARPin-PEG5K r = 3 nm, neutral) are constructed per established bioengineering [GROUNDED techniques: Gradisar-Jerala 2014 JACS for GCN4 trimers; Kim-Tezcan 2017 for ferritin-stable variants; Binz-Pluckthun 2004 for DARPin]. If nu_SE measured with these three scaffolds at the same nominal size differs by > 0.2 at any t_age, scaffold chemistry is a dominant source of variance and the probe-size scaling interpretation is moderated.
Bisociation: Polymer gel physics (inanimate entangled networks with controlled mesh contraction) <-> ALS protein condensates (biological mesh contraction via aberrant LCD-LCD interaction). The bridge is the SCALE-DEPENDENT PROBE MOBILITY: a universal consequence of any crosslinked/entangled meshwork, independent of the chemistry of crosslinking. Physics gave the exponent machinery; cell biology provides the pathological substrate.
Supporting Evidence
Stage 1a (in vitro xi_c measurement, PRIMARY prerequisite test):
Reconstituted TDP-43 condensates (200 uM, 150 mM NaCl, 20 mg/ml PEG-8000, 25 C). HaloTag-JF549-RRM1(F147L,F149L) monomer probe at 100 pM. nsFCS on condensate interior: fit G(tau) at 10 ns - 10 us with 2-component model. Extract xi_c = sqrt(D_fast * tau_crossover).
Primary prediction 1a: xi_c in TDP-43 WT at 1 h aging is in range 2-20 nm [PARAMETRIC, but bounded by Galvanetto 2023 precedent for generic coacervates]. Null: xi_c > 50 nm (requires larger probe panel and is out of FCS reach) or xi_c < 1 nm (FCS photon-statistics limit, D too fast to measure).
Stage 1b (scaffold-chemistry control, new):
Same-size 3 nm probes (GCN4-trimer basic, ferritin-trimer-stable acidic, DARPin-PEG5K neutral), each labeled with JF646 at 100 pM, each imaged separately in reconstituted TDP-43 WT condensate at t_age = 1 h.
Primary prediction 1b: D values across the three same-size scaffolds agree within 2x. Null: D values differ by > 3x -> scaffold chemistry dominates and size-scaling interpretation is invalid.
Stage 2 (3-probe size panel):
Probes: HaloTag-RRM1(F147L,F149L) monomer (r = 2.4 nm), same scaffold GCN4-trimer (r = 3.5 nm), ferritin-24mer (r = 12 nm). Measure D at t_age = 1, 4, 12, 24 h in TDP-43 WT and TDP-43 M337V (Patel 2015 faster-aging mutant, PMID 26317470 GROUNDED). K_p(r) measured at each time point via ratiometric confocal.
Primary prediction Stage 2: At t_age_optimal (4 h for M337V, 12 h for WT, hypothesized), nu_SE_corrected peaks at 0.5 - 1.0, with 95% CI excluding nu_SE < 0.3 for M337V and 95% CI excluding nu_SE > 0.5 for WT at t_age = 4 h. Null: nu_SE_corrected < 0.3 at all time points for both WT and M337V -> approach fails.
Stage 2b (dual-technique cross-validation, new for Directive 5):
Passive microrheology of 50-nm carboxylated polystyrene beads in same condensate aliquots. Extract G\*(omega) via GSER (Mason 2000, PMID 11033341 [GROUNDED topic + author + year]). Compute Maxwell time tau_M = eta/G'.
Primary prediction Stage 2b: In aged TDP-43-M337V (t_age = 4 h), where nu_SE_corrected > 0.7 is predicted, G(omega) should cross over from viscous-dominant at low omega to elastic-dominant at high omega with crossover frequency omega_c corresponding to 1/tau_M. If nu_SE > 0.7 but G(omega) remains purely viscous (G'' >> G' across all accessible omega), the mesh interpretation is falsified and alternative (e.g., compositionally distinct dense phase with higher bulk eta but no mesh) must be invoked.
Stage 3 (live-cell, conditional on Stage 2 Cohen's d > 0.8):
n = 3 per group: TDP-43 M337V patient iPSC-MN + 3 isogenic-corrected controls + 3 age-matched WT donors. Arsenite 200 uM, 30 min. 2-probe FCS (r = 3 nm, r = 12 nm). K_p(r) calibrated per cell.
Primary prediction Stage 3: D_small/D_large ratio is higher in M337V vs isogenic-corrected by > 2x, computed as median across stress granules within each donor.
- nu_SE > 1 is PARAMETRIC - direct polymer-gel measurements (Pluen 1999) bound nu_SE at approximately 1 in practical regimes. Claims of nu_SE > 1 require independent validation via G*(omega) showing entanglement plateau. Cycle 0 H1's nu_SE ~ 1.5 was a misattribution; cycle 2 version predicts nu_SE peaks at 0.5-1.0.
- Scaffold-chemistry confound - now explicitly tested via Stage 1b; if confound dominates, Stages 2-3 are moderated.
- K_p(r) partitioning bias - algebraic deconvolution formula: nu_SE_corrected = nu_SE_observed - (1/log(r_ratio)) * log(K_p_small/K_p_large). Measured at each t_age.
- TDP-43 fiber formation at long aging (Patel 2015) - ThT fluorescence parallel; exclude time points where ThT > 20% max.
- FCS photon-statistics floor for small fast probes - if D > 100 um^2/s, correlation too fast for standard FCS; require SP-FCS or STED-FCS.
- Stage 2b technique cross-check may disagree - this is the genuine negative result; the hypothesis is bounded by the probe-size-scaling vs G*(omega) convergence requirement.
How to Test
(1) Stage 1a (xi_c): Reconstitute TDP-43-WT-HaloTag condensate. 100-pM JF549-HaloTag-RRM1(F147L/F149L) probe. nsFCS 60 s per condensate, 10 condensates. Fit 2-component G(tau); extract xi_c per condensate.
(2) Stage 1b (scaffold chemistry): Three same-size-different-chemistry 3-nm probes (GCN4-basic, ferritin-acidic, DARPin-neutral); each at 10 pM in separate condensate aliquots; same TDP-43-WT base. FCS at t_age = 1 h. Compare D across scaffolds.
(3) Stage 2 (size panel): Three sizes (2.4, 3.5, 12 nm), each with GCN4-matched scaffold chemistry (same scaffold, different multimerization states). K_p(r) ratiometric confocal per probe per time point. FCS per probe per time point. Compute nu_SE_corrected = -[log(D_r2) - log(D_r1)] / [log(r2/r1)] - (1/log(r_ratio)) * log(K_p(r1)/K_p(r2)). Repeat for M337V.
(4) Stage 2b (dual-technique): 50-nm carboxylated polystyrene beads (0.01% v/v added at condensate formation). Passive microrheology video 100 Hz, 10 min per condensate. GSER -> G*(omega). Extract tau_M = 1/omega_c. Compare tau_M consistency with predicted from size-scaling nu_SE.
(5) Go/no-go: If Stage 2 nu_SE_M337V - nu_SE_WT > 0.3 at t_age = 4 h (n=5 replicates per condition, Cohen's d > 0.8), proceed to Stage 3. If Stage 2b G*(omega) disagrees qualitatively with Stage 2 size-scaling nu_SE, moderate conclusions and investigate scaffold-chemistry or probe-mesh interaction artifacts.
(6) Stage 3 (conditional live-cell): 3 M337V + 3 isogenic + 3 WT iPSC-MN donors. Arsenite stress, 2-probe FCS (the Stage-2-validated pair). K_p(r) per cell. Primary endpoint: D_small/D_large ratio across donors (mixed-model ANOVA with donor as random effect).
(7) Statistical test: Paired Wilcoxon signed-rank on nu_SE_corrected (Stage 2); mixed-model ANOVA for Stage 3. Primary alpha = 0.01 with Bonferroni for 4 time points.
Cross-Model Validation
Independent AssessmentIndependently assessed by GPT-5.5 Pro and Gemini Deep Research Max for triangulation. Assessed independently by two external models for triangulation.
Other hypotheses in this cluster
Maxwell Relaxation Time Aging Exponent beta_M in FUS-P525L Condensates
Tracking how fast diseased protein droplets 'solidify' could reveal a hidden clock in ALS progression.
Mutual Information I(X;Y) as Model-Free Liquidity Metric for Condensate State
Measuring how 'liquid' a cell's droplets are by tracking whether molecules move in sync — no physics model required.
Related hypotheses
Ferritin Protein Shell as Kinetic Barrier Controlling Ferrihydrite Fenton Activity
The protein cage surrounding your cells' iron stores may be a safety vault keeping a potent chemical reactor under lock and key.
Gaussian Mixture Model Analysis of Cryo-EM OMV Populations Distinguishes Biogenesis Pathways in P. aeruginosa
AI-powered microscopy could reveal how bacteria decide what to pack into their tiny 'mail packages'.
Abiotic vs Enzymatic PLOOH Regioselectivity as Chemical Fossil of Antioxidant Evolution
The chaotic chemistry of ancient iron reactions may have driven evolution of the precise enzymes that now control cell death.
Can you test this?
This hypothesis needs real scientists to validate or invalidate it. Both outcomes advance science.