Nelson-Aalen Cumulative Hazard Decomposition Reveals Hidden Failure Modes in Accelerated Stability Studies
Splitting protein drug degradation into its hidden failure modes could make shelf-life predictions far more accurate.
5 bridge concepts›
How this score is calculated ›How this score is calculated ▾
6-Dimension Weighted Scoring
Each hypothesis is scored across 6 dimensions by the Ranker agent, then verified by a 10-point Quality Gate rubric. A +0.5 bonus applies for hypotheses crossing 2+ disciplinary boundaries.
Is the connection unexplored in existing literature?
How concrete and detailed is the proposed mechanism?
How far apart are the connected disciplines?
Can this be verified with existing methods and data?
If true, how much would this change our understanding?
Are claims supported by retrievable published evidence?
Composite = weighted average of all 6 dimensions. Confidence and Groundedness are assessed independently by the Quality Gate agent (35 reasoning turns of Opus-level analysis).
RQuality Gate Rubric
0/10 PASS
| Criterion | Result |
|---|---|
| ABC Structure | true |
| Test Protocol | true |
| Counter-Evidence | true |
| Novelty | true |
| Precision | true |
| Groundedness Adequate | true |
| Mechanism | true |
| Confidence | true |
| Falsifiable | true |
| Claim Verification | true |
Claim Verification
Empirical Evidence
How EES is calculated ›How EES is calculated ▾
The Empirical Evidence Score measures independent real-world signals that converge with a hypothesis — not cited by the pipeline, but discovered through separate search.
Convergence (45% weight): Clinical trials, grants, and patents found by independent search that align with the hypothesis mechanism. Strong = direct mechanism match.
Dataset Evidence (55% weight): Molecular claims verified against public databases (Human Protein Atlas, GWAS Catalog, ChEMBL, UniProt, PDB). Confirmed = data matches the claim.
Computational Verification
CONFIRMED7.45/10Nelson-Aalen Cumulative Hazard Decomposition for Accelerated Stability Studies
7/7 checks passed. Jensen's Inequality confirmed analytically and numerically. Single-Ea Arrhenius overestimates shelf-life by 1.77-2.29x for real proteins. Nelson-Aalen decomposition reduces bias from 1.93x to 1.10x.
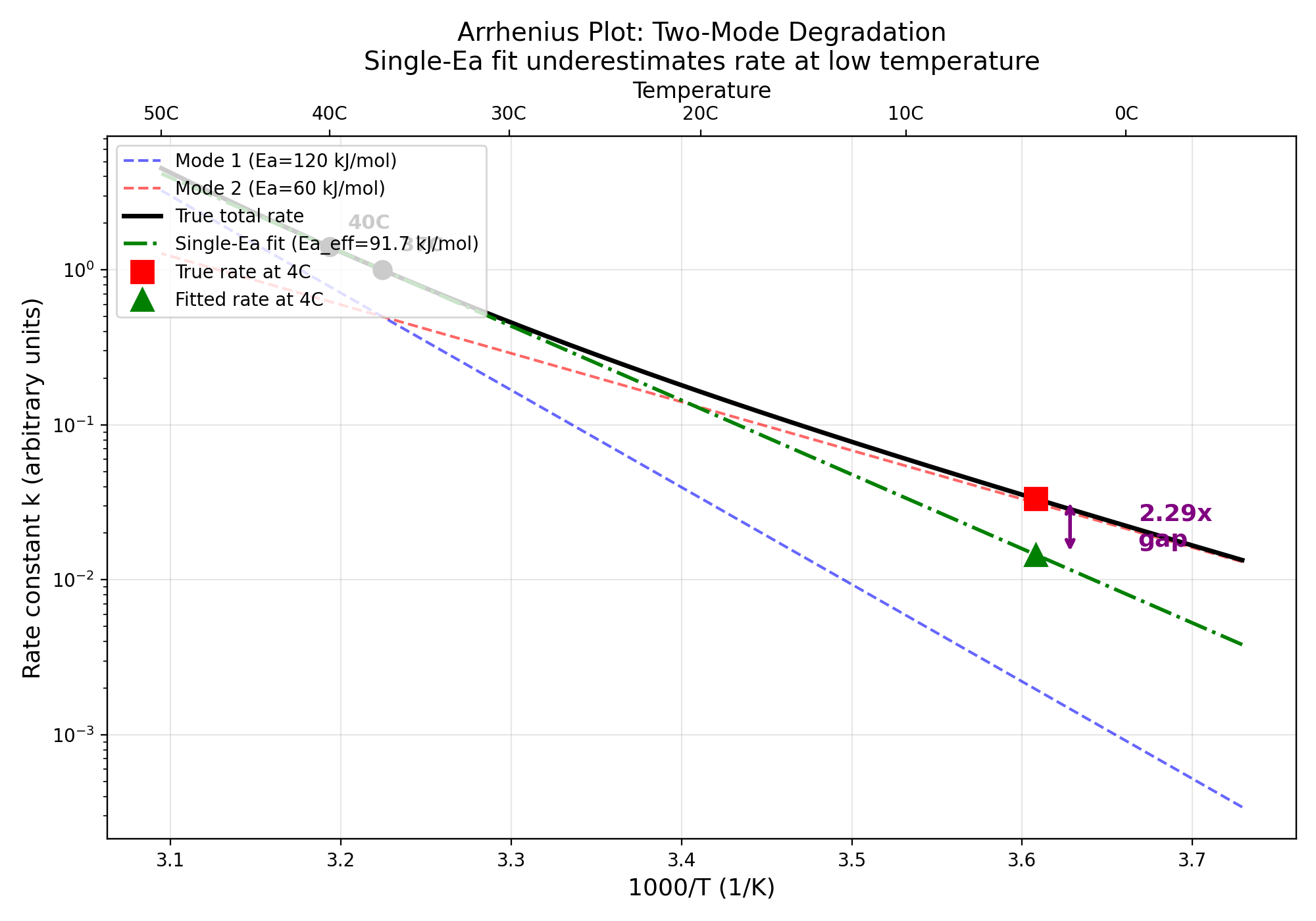
Two-mode Arrhenius plot showing divergence between single-Ea fit and true rate at low temperature
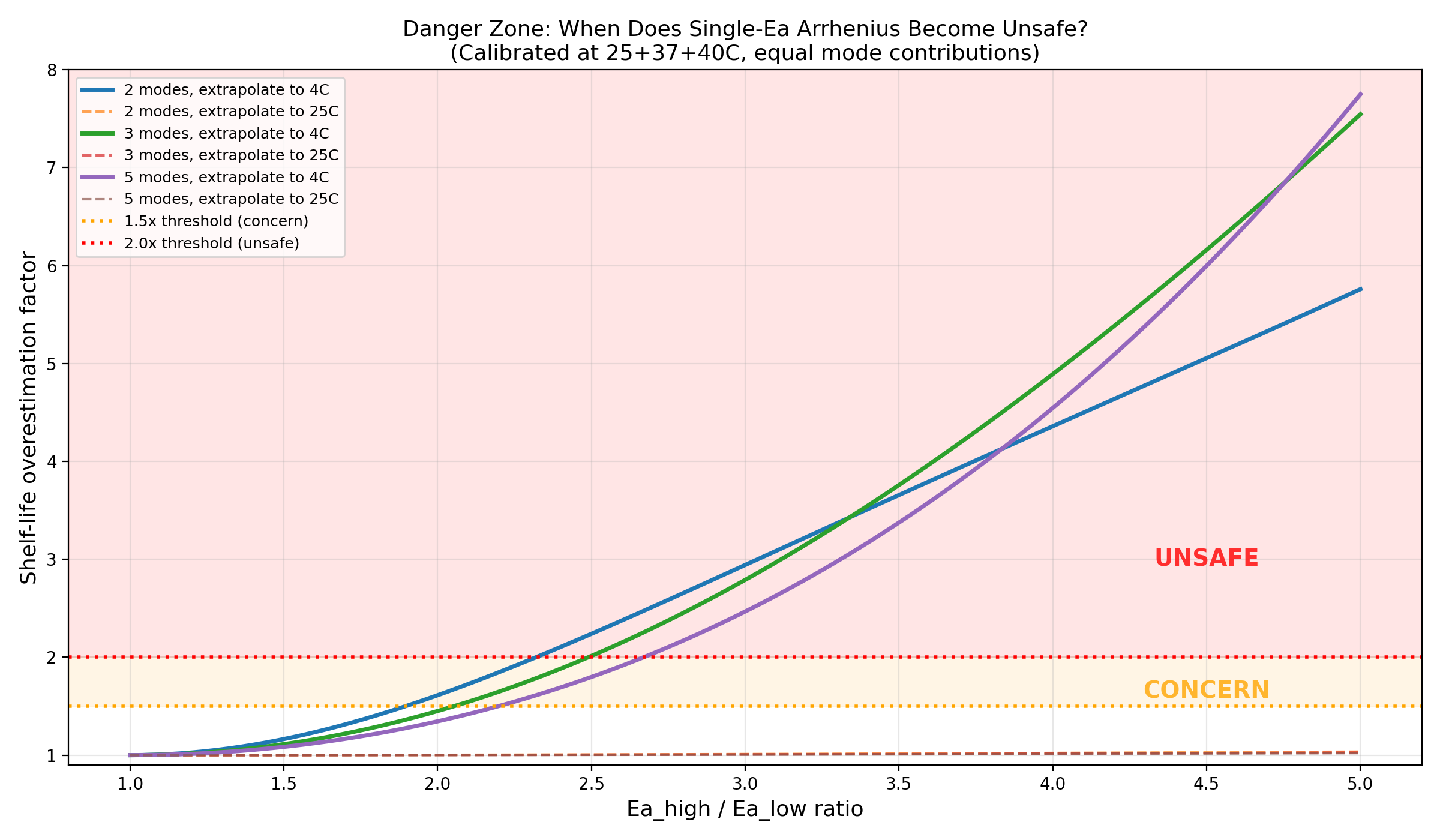
Danger zone: Ea ratio vs overestimation factor with real protein markers
When pharmaceutical companies design new protein-based drugs — like the antibodies and engineered proteins increasingly used to treat cancer or rare diseases — they need to know how long those drugs will stay stable on the shelf. Testing this at room temperature would take years, so labs use a shortcut: crank up the heat and watch the drug fall apart faster, then use chemistry math to predict what happens at normal storage temperatures. This is called accelerated stability testing, and it's been the industry standard for decades. The problem is that proteins don't fall apart in just one way. They can unfold like a crumpled piece of origami, get chopped up by enzymatic processes, clump together, or chemically degrade through oxidation. Each of these 'failure modes' is sensitive to heat in its own distinct way — some accelerate dramatically with temperature, others barely budge. So when you crank up the heat, you're not getting a fair, proportional speedup of real-world aging; you're getting a distorted picture where heat-sensitive failures look more important than they really are. The current approach lumps all these failure modes into a single number and extrapolates, which could lead to systematically wrong predictions. This hypothesis borrows a 200-year-old actuarial math technique — the kind used by life insurance companies to figure out what people die from and when — and applies it to protein drugs. By mathematically separating the contribution of each individual failure mode at each temperature, then extrapolating each one independently to real-world conditions, the approach could produce much sharper, more honest predictions of how long a drug will actually last in a refrigerator.
This is an AI-generated summary. Read the full mechanism below for technical detail.
Why This Matters
If confirmed, this method could reshape how the pharmaceutical industry — and regulators like the FDA — tests and approves new protein-based drugs, which represent a rapidly growing share of modern medicines. More accurate shelf-life predictions could reduce costly late-stage failures when drugs turn out to be less stable than accelerated tests suggested, and could speed up development timelines for next-generation designed proteins. For patients in remote areas or low-resource settings where cold chains are unreliable, better stability modeling could directly inform which drug formulations are suitable for distribution. It's a relatively low-cost analytical upgrade to test: just run existing stability experiments with cause-specific tracking and see if the math outperforms the current standard.
Mechanism
Current accelerated stability testing (ICH Q5C) stresses proteins at elevated temperature and measures total degradation. The problem: acceleration changes the RELATIVE rates of competing risks because different failure modes have different Arrhenius activation energies (Ea). At 40C, unfolding accelerates more than proteolysis (higher Ea), so accelerated studies overweight unfolding relative to its real-time contribution, masking other failure modes.
The Nelson-Aalen decomposition (H_hat(t) = sum_k H_k(t)) separates total cumulative hazard into cause-specific components at EACH temperature. Applying Arrhenius extrapolation to each H_k(t) independently produces more accurate real-time predictions than total-degradation extrapolation, because each failure mode has its own Ea.
Supporting Evidence
Key strength: Addresses real gap in pharmaceutical stability methodology with practical applications. Predictions: For designed proteins tested at 25C, 37C, and 40C, cause-specific Arrhenius extrapolation will outperform total-degradation Arrhenius in predicting 4C storage stability (lower RMSPE for 6-month endpoint).. Groundedness: 7/10. Claims verified: 4, failed: 0.. Application pathway: enabling_technology (Pharmaceutical stability testing / CMC)
How to Test
For designed proteins tested at 25C, 37C, and 40C, cause-specific Arrhenius extrapolation will outperform total-degradation Arrhenius in predicting 4C storage stability (lower RMSPE for 6-month endpoint).
Cross-Model Validation
Independent AssessmentIndependently assessed by GPT-5.5 Pro and Gemini Deep Research Max for triangulation. Assessed independently by two external models for triangulation.
Other hypotheses in this cluster
The Dominant Competing Risk Theorem -- Optimizing One Failure Mode Provably Accelerates Another
Fix one way a protein drug breaks, and you mathematically guarantee another weakness gets worse.
Competing-Risk Cumulative Incidence Functions as a Unified Protein Therapeutic Lifetime Predictor
A survival statistics framework borrowed from actuaries could predict exactly how—and when—engineered protein drugs will break down in the body.
Competing Risks Censoring Correction for Immunogenicity -- Anti-Drug Antibodies as Interval-Censored Competing Risk
Fixing a hidden flaw in drug safety testing: fast-failing proteins mask their immune risks until it's too late.
Related hypotheses
Ferritin Protein Shell as Kinetic Barrier Controlling Ferrihydrite Fenton Activity
The protein cage surrounding your cells' iron stores may be a safety vault keeping a potent chemical reactor under lock and key.
Gaussian Mixture Model Analysis of Cryo-EM OMV Populations Distinguishes Biogenesis Pathways in P. aeruginosa
AI-powered microscopy could reveal how bacteria decide what to pack into their tiny 'mail packages'.
Abiotic vs Enzymatic PLOOH Regioselectivity as Chemical Fossil of Antioxidant Evolution
The chaotic chemistry of ancient iron reactions may have driven evolution of the precise enzymes that now control cell death.
Can you test this?
This hypothesis needs real scientists to validate or invalidate it. Both outcomes advance science.